Files
Download Full Text (1.5 MB)
Publication Date
5-2023
Abstract
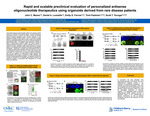
The generation of patient derived iPSCs has revolutionized the field of personalized medicine. With the unlimited capacity to self-renew and the ability to differentiate into any cell type, iPSCs hold great promise to develop cell based personalized therapies for rare diseases. Antisense oligonucleotides (ASOs) have emerged as a promising therapeutic approach for the treatment of various diseases; and several ASOs for rare diseases have received clinical approval, confirming the potential of this approach. With the advancement of next generation sequencing technologies, the ability to identify disease variants that could benefit from ASO therapeutics has grown. To take advantage of these technologies, we have developed a platform to expediate the rapid characterization of preclinical ASO leads using our iPSC pipeline. We developed an efficient and scalable iPSC pipeline that allows for the rapid generation of iPSCs from patient PBMCs that can be completed in 4-6 weeks. Our reprogramming success rate is >90%, where we can simultaneously generate dozens of iPSCs in parallel, allowing for the generation of >100 patient derived iPSCs in less than 3 months. Using Genomics Answers for Kids (GA4K), we identified a panel of Duchenne muscular dystrophy (DMD) patients that were amendable to ASO therapy. We generated patient derived iPSCs from the DMD panel. In this panel, one patient harbored a structural deletion of exons 46-53 of the DMD gene making it amendable to treatment with the FDA-approved ASO Casimersen. Patient-derived iPSCs treated with ASOs matching the Casimersen sequence showed restoration of Dystrophin protein within 5 days. The DMD panel also included a pair of siblings, both harboring a deep intronic variant in the DMD gene that gives rise to a novel splice acceptor site, incorporation of a cryptic exon, and premature transcript termination. We designed ASOs targeting the intronic variant and observed restoration of Dystrophin protein expression in the patient’s iPSC lines. Next, we differentiated the patient iPSC lines into cardiac organoids and examined organoid function. The cardiac organoids from DMD patients displayed weak arrhythmic contractions and abnormal fluctuations in calcium levels. Treatment with patient specific ASOs before organoid differentiation or post-differentiation, restored both contraction rates and calcium levels comparable to control. Overall, we were able to obtain patient samples, generate patient derived iPSCs, and validate personalized ASOs in fewer than 8 weeks. This platform can be adapted to other cellular models, such as brain organoids, allowing for a rapid and scalable preclinical evaluation of personalized ASOs.
Document Type
Poster
Recommended Citation
Means, John C.; Louiselle, Daniel A.; Koseva, Boryana; Pastinen, T; and Younger, Scott T., "Rapid and scalable preclinical evaluation of personalized antisense oligonucleotides using organoids derived from rare disease patients" (2023). Research at Children's Mercy Month 2023. 14.
https://scholarlyexchange.childrensmercy.org/research_month2023/14