Files
Download Full Text (627 KB)
Publication Date
5-2024
Abstract
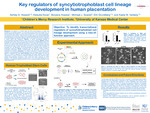
In human placentation, cytotrophoblast cells fuse to form a continuous epithelial cell layer of syncytiotrophoblast (ST). Multinucleated ST cover the villus surface and provide critical structural and biochemical barriers for the developing fetus. ST also produce hormones and growth factors to support pregnancy and promote fetal growth. Despite these critical functions, key transcription factors (TF) and signaling pathways involved in ST differentiation are not fully understood. To identify regulators of ST differentiation, we integrated transcriptomic and genome-wide epigenomic approaches in human trophoblast stem (TS) cells. Human TS cells were maintained in a proliferative stem state or differentiated into ST following a six-day ST3D differentiation protocol. Stem and ST differentiated cells were analyzed using RNA-sequencing, Assay for Transposase-Accessible Chromatin-sequencing, and HiC chromatin capture. Prominent differences in chromatin accessibility, cell transcriptomes and TF binding motifs were identified in Stem and ST cell states. The top ten TF binding motifs identified in ST-specific regions were enriched for AP-1 family members (e.g., Fra1, Fos, JunB, Fra2, BATF, Atf3, Fosl2, AP-1, Jun-AP1), TEAD, and AP-2gamma. Select TFs identified as uniquely enriched in ST cells were localized in human placental tissues by in situ hybridization and mechanistically studied using loss-of-function approaches in the TS cell model. Overall, key regulators of syncytiotophoblast cell lineage development in human placentation were identified.
Document Type
Poster
Recommended Citation
Howard, Ashley; Kozai, Keisuke; Koseva, Boryana; Soares, Michael J.; Grundberg, Elin; and Varberg, Kaela, "Key regulators of syncytiotrophoblast cell lineage development in human placentation" (2024). Research Month 2024. 3.
https://scholarlyexchange.childrensmercy.org/research_month2024/3